
![]()



![]()
![]()
 |
As a graduate student and
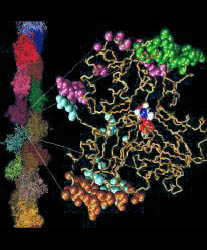
postdoc in the 1990s I was fascinated by the newly solved structures of G- and F-actin. I performed
some of the earliest Molecular Dynamics (MD) simulations of actin and of the
motor protein kinesin and collaborated with various experimental groups
in the motility field. The work also led to significant method
development, notably in protein domain motion analysis.
|
 |
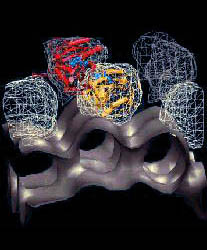
In 1998 I came across a research problem that dramatically accelerated my
academic career. I developed a coarse-grained model and used it for bringing
G-actin structures into register with F-actin electron microscopy (EM)
maps that were rendered at the same level of detail [see paper]. This work led to the development of the award-winning Situs
package, opened the door to a faculty position at The Scripps Research
Institute, and it created opportunities for rigid body and flexible
fitting of structural data from various biophysical origins.
|
 |
The integration of
multi-scale data from various biophysical origins soon became
our signature research area and earned us an NIH R01 grant, due to the need in the "low
resolution" community to produce atomic models for their data.
Correlation techniques had arguably the biggest impact in this
approach. In 2001 Pablo
Chacón invented
the Laplacian filter and implemented Fast Translational Matching in the
now-famous colores
program. Julio
Kovacs and Yao Cong later
developed the Fast Rotational Matching approach in 2D, 3D, and 6D which
earned us a HFSP
project grant.
|
 |
Haptic devices and
virtual reality became a research focus after 2001. Force
feedback during interactive EM fitting in our
3D
theater
was an amazing experience for the user. The focus in later years turned to improving the graphics
and functional capabilities of the Sculptor
program developed in the lab. Our work was far ahead of what much bigger
groups and consortia were developing at the time.
|
 |
During my work on actin in
the 1990s I met the inventor of elastic network models, Monique Tirion.
In 2002 we applied ENMs for
the first time to EM maps in collaboration / competition with Florence
Tama and Charles Brooks. Later, some of our ideas led to the ModeHunter package which is still
actively developed.
|
 |
Large scale MD simulations produce an immense
quantity of data.
Principal
Component Analysis (PCA) is a standard mathematical tool used to detect
correlations in large data sets, but one problem we encountered in the
1990s was that the PCA modes suffer from the MD sampling problem.
Zhiyong Zhang and I therefore developed a localized set of basis
functions to
discriminate relevant conformational changes in a protein from the
background of atomic fluctuations. More recently, I have adapted
computational geometry and network techniques to the study
of long-time scale trajectories generated at D.E. Shaw Research.
|
![]()